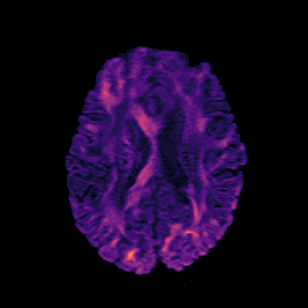
Patch2Self improves the speed of clinical MRI while also improving the quality of diffusion MRIs and all dMRI analysis.
Researchers from the Department of Intelligent Systems Engineering at the Luddy School of Informatics, Computing, and Engineering have developed a method for recovering the signal and automatically denoising diffusion magnetic resonance imaging data.
The method, Patch2Self, improves the speed of clinical MRI while also improving the quality of diffusion MRIs and all dMRI analysis. Patch2Self doesn’t require external datasets or prior information to perform denoising, allowing it to be used with current MRI data.
“By removing the noise, we increase the conspicuity of the image,” said Eleftherios Garyfallidis, an assistant professor of intelligent systems engineering at the Luddy School and the director of the Garyfallidis Research Group. “Then, the images are easier to interpret by the doctor and correspond more accurately to the underlying anatomy. Furthermore, all forthcoming analyses become more robust. It is well known that noise can impede any signal processing mechanism. By suppressing the noise, we directly reduce confounding factors in the acquired signal, which leads to better analysis; for example, reducing false positives in detecting pathology. At GRG, we are pioneering a new generation of tools to address these issues using machine learning, ensuring speed, accuracy and clinical viability of the proposed solutions.”
Diffusion MRI is one of the most commonly used medical imaging technologies, and quick and accurate results are critical to the wellbeing of patients. Higher resolution images are ideal, but current diffusion MRI techniques create noisy images. Patch2Self uses self-supervised learning and the statistical independence of noise to suppress confounding signals in MRI images, resulting in higher resolution images.
“Patch2Self’s novelty lies in the idea that if one had four-dimensional data, you could build an optimal denoising function by using a part of the data as a noisy signal to predict itself,” said Shreyas Fadnavis, a Ph.D. candidate in ISE who is the lead author of the study. “Since the noise is uncorrelated, the proposed algorithm ends up learning the true underlying signal. This essentially paves way for suppressing different types of noise induced in the MR signal acquisition.”
Fadnavis recently presented his work at the Neural Information Processing Systems Conference in December. The paper also received a spotlight presentation at the premier event for machine learning researchers.
“The proposed method is self-supervised and has no hyperparameters,” Garyfallidis said. “This means that it is a fully automated method. This is remarkably uncommon and an important landmark for machine learning applied to medical imaging. This is one of the key goals of machine learning to build such battle-tested systems that need no work on the operator's/user's side. Usually, many techniques still require a lot of manual control. In addition to improving our capacity in using self-supervision in medical imaging, Patch2Self opened new, unexpected research horizons into studying theoretical principles of diffusion MRI signal.”
Patch2Self, which was developed in close collaboration with Joshua Batson from Chan Zuckerberg Biohub, has been widely disseminated to the community via DIPY as a thoroughly tested and data agnostic framework.
“Researchers at the Luddy School are consistently pushing the boundaries of what is possible,” said Kay Connelly, the associate dean for research at the Luddy School. “The work of Shreyas, Eleftherios, and the Garyfallidis Research Group is creating new pathways to improving digital imaging, and this critical advancement will create new opportunities to make a real-world impact.”

